Research in the centre focuses on structural and functional studies of the main building blocks of living organisms – proteins, DNA, membranes, solvents and small molecules such as drug compounds. We develop cutting-edge, state-of-the-art techniques to model molecular phenomena taking place at the intracellular level, and our software applications are some of the most widely used in the world in our domain. We apply the techniques and methods to address some of the biggest scientific challenges in the area of biomolecular research, as described below.
Antibody therapeutics
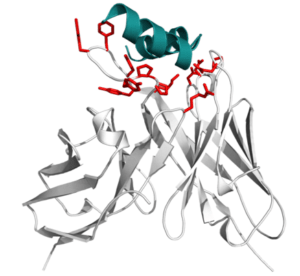
Many diseases still lack treatment with conventional drugs. In recent years, antibodies have been developed as therapeutics in the fight against diseases, due to their lower side effects compared with traditional chemotherapy. An antibody drug generally works by attaching itself to an antigen, which is a unique site of the pathogen and harnessing the immune system to directly attack and destroy it. Currently approved treatments with antibodies target many diseases including some types of cancer. However, antibody development is an expensive and labour-intensive process. Within BioExcel, we use computational tools to study the interactions between antibodies and antigens. In this process, the most promising drug candidates are selected, and workflows are applied to simulate how these drugs candidates interact with the target. Through this project, our ultimate goal is to shorten the overall time and effort taken to develop new therapeutics for patients.
Modelling the human proteome
The human proteome consists of over 20000 proteins, which are the molecular machines responsible for the most important cellular processes. Most of these machines work by interacting with each other and with other biomolecules present in our cells. This highly intricate and complex network of interactions is called the interactome. These interactions regulate life and malfunctions, induced either by the enviroment or via natural mutations, are responsible for many diseases. Understanding how these proteins interact in a detailed level is crucial for the development and improvment of new drugs and treatments. However the computational analysis and prediction of this web of interactions is a highly demaning task. Adding the atomic information to thousands of protein complexes requires a robust, high perfomance computational infrastructure.

Rational Drug Design
Rational drug design refers to computationally designing drug-like molecules that bind to a macromolecule. It is routinely used in industry to guide the synthesis and experimental testing of preclinical molecules towards a drug candidate. We aim at delivering workflows that automate and accelerate a broad range of drug design tasks. Using structural knowledge and molecular dynamics-based techniques such as pmx/GROMACS, we assess the effects of mutations on binding. We are developing four workflows of increasing complexity to (i) identify mutations causing drug resistance, (ii) repurpose drugs for recovering protein function, (iii) optimize peptides for better protein binding, and (iv) predict binding “strengths” (affinities) via machine learning based on molecular dynamics data.

Protein Dynamics
Mass spectrometry has revolutionized proteomics, i.e. the investigation of the myriad of protein/protein interactions in the cell. By using a very small sample content of proteins (usually in the micromolar concentration), a powerful implementation of the technique, the so-called electrospray ionization/ion mobility mass spectrometry (ESI/IM-MS), can measure stoichiometry, shape and subunit architecture of protein and protein complexes in vacuum. The key assumption in the ESI/IM-MS technique is that the rearrangement of biomolecular units on passing from solution (i.e. in the cell) to vacuum is minimal. However, proton transfer phenomena that could happen in vacuum and not in solution may significantly affect structural properties. Therefore, understanding the mechanisms and quantifying the extent of these configurational changes in vacuum will allow significant improvement in the predictions of this already powerful experimental technique when exporting this information to the cell environment. In particular, within BioExcel we use massively parallel simulations to address this problem on an important example use case system, the protein beta-lactoglobulin. Comparison with available experimental data will allow us to establish the reliability and accuracy of ESI/IM-MS measurements in describing structural features of proteins in cells.

Fluorescence microscopy
Fluorescence microscopy with fluorescent proteins is an essential technique for revealing how cells function and for developing new strategies to fight diseases. After fusing a fluorescent protein to another protein using gene technology, the exact location of that protein can be determined by monitoring the fluorescence emitted by the photo-excited fluorescent protein with a normal microscope. The resolution of this technique is determined by the properties of the fluorescent proteins, rather than lenses. However, these proteins still require significant improvements of their properties to unlock the full potential of this technique for biological imaging. To overcome this limitation, we develop a modelling protocol that performs atomistic simulations based on a validated physical models. From the simulations, all relevant properties of the protein are extracted, such as protein stability, absorption and fluorescence spectra, as well as quantum yields.
More information about current and previous projects can be found here.
Related scientific areas
It is important to emphasize that our area of expertise, i.e. biomolecular modelling:
- Is the central computational technique used in biochemistry to determine and improve structure.
- Is the most important tool for high-throughput virtual screening in pharmaceutical drug design.
- Is one of the most computationally intensive part of life science research (in terms of core hours)
- Lays the foundations for understanding higher levels of organization of biological systems.
- Is applicable across a broad range of research including non-biological studies of soft matter.
Thus, BioExcel CoE activities support researcher work in other core disciplines in the computational sciences such as:
- Integrative structural biology
- Biomarkers design
- Nanotechnology and materials science
- Personalized medicine
- Physiology
- Neuroinformatics
- Multi-scale QM/MM modelling
Here is an overview of the role BioExcel can play in those disciplines.
